More complex machines are more likely to break - they offer more potential breaking points. Yet, cells are highly complex entities and are extremely robust, i.e. they perform their functions reliably even in variable intra- and extracellular milieus. We want to understand the underlying basis: what makes biological systems robust?
To study this question, we look at cell growth and division. While cells must have emerged from something simpler billions of years ago, today the only way we know of for a new cell to emerge is by division from another cells. Cell growth and division therefore are the basis of life, and their deregulation is common in cancer. We want to know: How do cells grow in a coordinated fashion? What informs when they divide? And how do cells orchestrate cell division?
Any error in these processes can be fatal. Hence, reliability and robustness are crucial. To understand the underlying molecular networks and their dynamic behavior, we combine molecular genetics with live-cell quantitative microscopy, and biochemical experiments.
While traditionally the focus of the field has been on the regulatory protein networks, we find it important to consider not only the proteins, but also the underlying gene expression; they are all part of one integrated system.
Computational modeling, for which we collaborate with experts in this field, helps us to interpret our experiments and to develop new hypotheses. We work with fission yeast, which is an excellent model for eukaryotic cells, and where we can easily introduce genetic modifications.
How are cell division processes appropriately timed and coordinated with each other?
The separation of the chromosomes in anaphase is one of the most dramatic events of the cell cycle. The separation is irreversible and therefore needs to be highly accurate and tightly coordinated with mitotic exit. We combine perturbation experiments, live cell imaging and computational modeling to understand how this high degree of coordination is achieved. We want to find general principles that allow cells to execute highly dynamic processes with great accuracy.
How do cells keep variation within an acceptable range?
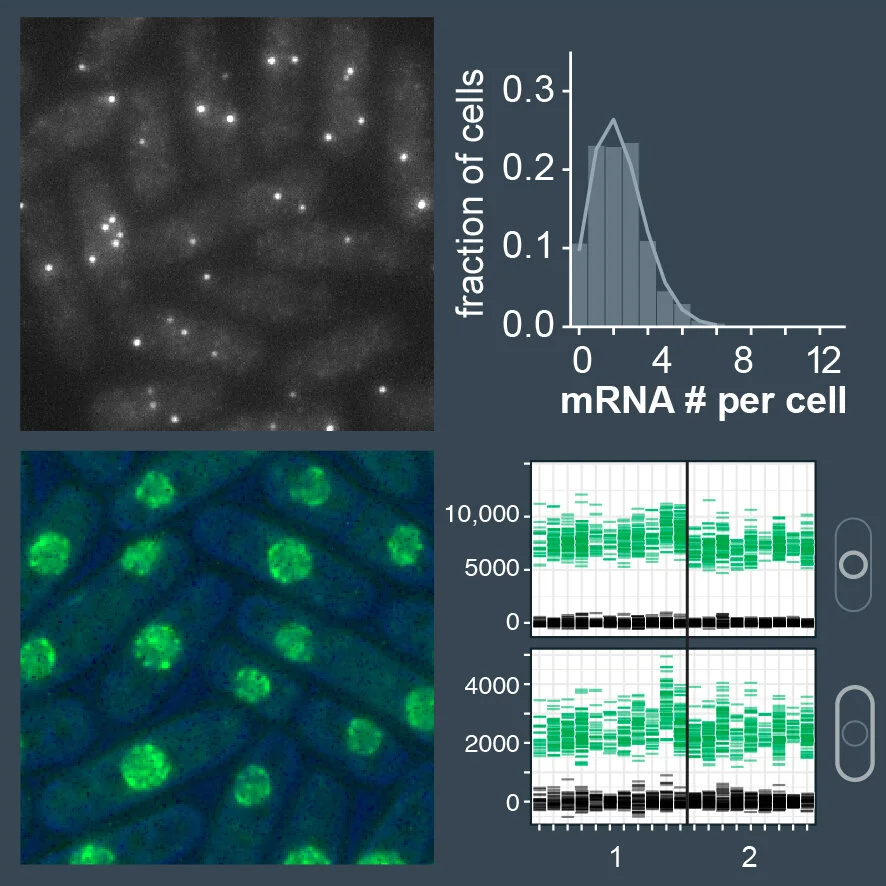
Biochemical reactions inside cells are fundamentally stochastic, i.e. a reaction occurs when two molecules encounter each other. If molecules are present in many copies, this happens frequently. However, in reactions that involve genomic DNA (1 or 2 copies per cell) or mRNA (often only a few copies per cell), this causes considerable variability and results in fluctuations of mRNA and protein abundance (called 'gene expression noise'). We want to know how cells keep noise in check and produce just the right amount of a given protein. We study this in the context of the spindle assembly checkpoint, where we know that precise protein concentrations are important for proper function.
Funding
We are extremely grateful for funding by the NIH and NSF!
Collaborators
Sahand Jamal Rahi / EPFL Lausanne / automated image analysis for fission yeast
Ramon Grima / University of Edinburgh / mathematical modeling / gene expression noise
Abhyudai Singh / University of Delaware / computational modeling / gene expression noise
Jing Chen / Systems Biology, Virginia Tech / computational modeling / gene expression
Andrea Ciliberto / IFOM, Milan, Italy / computational modeling / spindle assembly checkpoint
Stefan Legewie, Alex Anyaegbunam / IMB, Mainz, Germany / computational modeling / regulation of anaphase
+ Past collaborators
Lenwood S. Heath / Department of Computer Science, Virginia Tech / bioinformatics / spindle assembly checkpoint
Nicole Radde, Eva-Maria Geissen / University of Stuttgart, Germany / computational modeling / spindle assembly checkpoint
Jan Hasenauer / Helmholtz Zentrum Muenchen, Germany / stochastic modeling and statistics / spindle assembly checkpoint
Boris Macek, Karsten Krug, Alejandro Carpy / Proteome Center, University of Tuebingen, Germany / mass spectrometry / proteome and phosphoproteome across the cell cycle
Michael Knop, Susanne Trautmann / EMBL and ZMBH Heidelberg / fluorescence correlation spectroscopy
Gunnar Raetsch, Chris Widmer, Philipp Drewe / Memorial Sloan-Kettering Cancer Center / image recognition
Daniel Rauh / Chemical Genomics Centre of the Max Planck Society and Technical University Dortmund / chemistry / analog-sensitive kinases